PyClone 是一种用于推断癌症中克隆种群结构的统计模型。 它是一种贝叶斯聚类方法,用于将深度测序的体细胞突变集分组到假定的克隆簇中,同时估计其细胞流行率(prevalences)并解释由于分段拷贝数变化(segmental copy-number changes)和正常细胞污染(normal-cell contamination)引起的等位基因失衡。 单细胞测序验证证明了 PyClone 的准确性。
The input data for PyClone consists of a set read counts from a deep sequencing experiment, the copy number of the genomic region containing the mutation and an estimate of tumour content.
简易安装
官方推荐使用 MiniConda 来安装 PyClone。为了保证环境的稳定,可为 PyClone 单独建立一个环境,因为 PyClone 基于 Python2.7。在这里,我们使用 Anaconda3(conda 4.5.11) 来安装 PyClone。
# 创建基于 Python2.7 名字为 pyclone 独立环境conda create --name pyclone python=2# 激活 pyclone 环境source activate pyclone# 退出 pyclone 环境source deactivate# 安装 PyCloneconda install pyclone -c aroth85
Anaconda3 中安装完 PyClone,激活环境后,执行 PyClone -h 出现 RuntimeWarning。同样的,我们在 pyclone 的环境中导入 pandas 模板,出现一样的 RuntimeWarning:
(pyclone) shenweiyan@ecs-steven 13:38:25 /home/shenweiyan$ PyClone -h/usr/local/software/anaconda3/envs/pyclone/lib/python2.7/site-packages/pandas/_libs/__init__.py:4: RuntimeWarning: numpy.dtype size changed, may indicate binary incompatibility. Expected 96, got 88from .tslib import iNaT, NaT, Timestamp, Timedelta, OutOfBoundsDatetime/usr/local/software/anaconda3/envs/pyclone/lib/python2.7/site-packages/pandas/__init__.py:26: RuntimeWarning: numpy.dtype size changed, may indicate binary incompatibility. Expected 96, got 88......from pandas._libs import algos, lib, writers as libwriters/usr/local/software/anaconda3/envs/pyclone/lib/python2.7/site-packages/statsmodels/nonparametric/kde.py:22: RuntimeWarning: numpy.dtype size changed, may indicate binary incompatibility. Expected 96, got 88from .linbin import fast_linbin/usr/local/software/anaconda3/envs/pyclone/lib/python2.7/site-packages/statsmodels/nonparametric/smoothers_lowess.py:11: RuntimeWarning: numpy.dtype size changed, may indicate binary incompatibility. Expected 96, got 88from ._smoothers_lowess import lowess as _lowessusage: PyClone [-h] [--version]{setup_analysis,run_analysis,run_analysis_pipeline,build_mutations_file,plot_clusters,plot_loci,build_table}...positional arguments:{setup_analysis,run_analysis,run_analysis_pipeline,build_mutations_file,plot_clusters,plot_loci,build_table}setup_analysis Setup a config file and mutations files for a PyCloneanalysis.run_analysis Run an MCMC sampler to sample from the posterior ofthe PyClone model.run_analysis_pipelineRun a full PyClone analysis.build_mutations_fileBuild a YAML format file with mutation data and statesprior to be used for PyClone analysis.plot_clusters Plot features of the clusters.plot_loci Plot features of the loci.build_table Build results table which contains cluster ids and(mean) cellular prevalence estimates.optional arguments:-h, --help show this help message and exit--version show program's version number and exit(pyclone) shenweiyan@ecs-steven 14:47:17 /home/shenweiyan$ pythonPython 2.7.15 | packaged by conda-forge | (default, Oct 12 2018, 14:10:50)[GCC 4.8.2 20140120 (Red Hat 4.8.2-15)] on linux2Type "help", "copyright", "credits" or "license" for more information.>>> >>> import pandas/usr/local/software/anaconda3/envs/pyclone/lib/python2.7/site-packages/pandas/_libs/__init__.py:4: RuntimeWarning: numpy.dtype size changed, may indicate binary incompatibility. Expected 96, got 88from .tslib import iNaT, NaT, Timestamp, Timedelta, OutOfBoundsDatetime/usr/local/software/anaconda3/envs/pyclone/lib/python2.7/site-packages/pandas/__init__.py:26: RuntimeWarning: numpy.dtype size changed, may indicate binary incompatibility. Expected 96, got 88from pandas._libs import (hashtable as _hashtable,....../usr/local/software/anaconda3/envs/pyclone/lib/python2.7/site-packages/pandas/io/pytables.py:50: RuntimeWarning: numpy.dtype size changed, may indicate binary incompatibility. Expected 96, got 88from pandas._libs import algos, lib, writers as libwriters>>> pandas.__version__u'0.23.4'
原因与解决:(参考 anaconda-issues:#6678、numpy issues:#11628)
The pandas were build agains different version of numpy. we need to rebuild pandas agains the local numpy.
# 方法一(耗时长)pip install --no-binary pandas -I pandas# 方法二(推荐使用)conda install numpy==1.14.5 --yes
手动安装
要手动安装 PyClone,请确保安装了必要的库(如下所列)。 之后就可以像任何其他 Python 包一样通过 python setup.py install 安装 PyClone。
PyClone 必须满足依赖包如下:
PyDP >= 0.2.3PyYAML >= 3.10matplotlib >= 1.2.0 - Required for plotting.numpy >= 1.6.2 - Required for plotting and clustering.pandas >= 0.11 - Required for multi sample plotting.scipy >= 0.11 - Required for plotting and clustering.seaborn >= 0.6.0
手动安装 PyClone:
$ git clone https://github.com/aroth85/pyclone.git$ cd pyclone$ python setup.py installrunning installrunning bdist_eggrunning egg_infocreating PyClone.egg-infowriting PyClone.egg-info/PKG-INFO......Installed /usr/local/software/python2.7/pyclone/lib/python2.7/site-packages/PyClone-0.13.1-py2.7.eggProcessing dependencies for PyClone==0.13.1Finished processing dependencies for PyClone==0.13.1
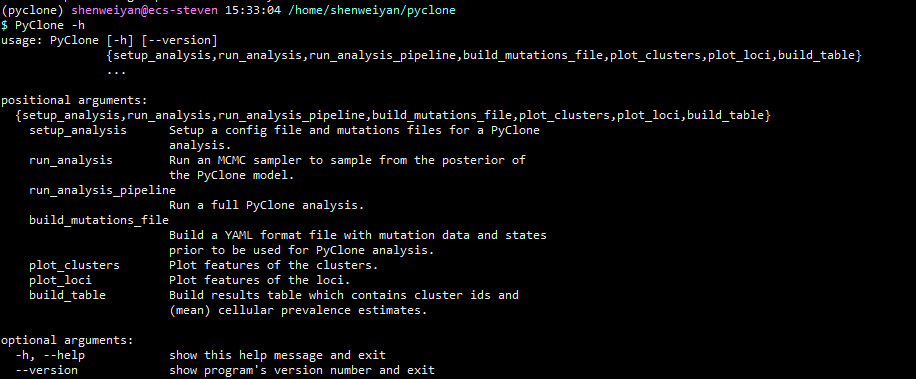
到这里,PyClone 就安装完成了,关于该软件具体的使用说明,请参考 PyClone -h 或者 PyClone wiki: Usage。
参考资料:
- numpy issues,#11628
- anaconda-issues,#6678
- aroth85/pyclone,GitHub
- YTer,Pyclone 说明,Hexo 个人博客
- 用户1680321,安装使用pyclone进行克隆演化推断,yw的数据分析

