- Result 1:Performance of different filtering parameters in Samtools fastq:
- (1)-f 3: (Output file size:312M)
- Choose the flag = 0x1 (paired-end) and the flag = 0x2 (each segment properly aligned according to the aligner) to include the reads needed.
- (2)-F 12: (Output file size:317M)
- Choose the flag = 4(segment unmapped) and the flag = 8(next segment in the template unmapped) to exclude the reads not needed.
- (3)-F 0x900: (Output file size:319M)
- Recommended by the help document in order to obtain the only paired reads by excluding the flag = 0x100 (secondary alignment)and the flag = 0x800 (supplementary alignment)
- (4)Parameter explainations:
- Flags explainations:**
- ">

- Result 2: Assess the assemblies with different subsampling coverage and SPAdes parameters (—meta/—isolate)
- Result 3: Package the pipeline :
- Result 4: Summary:
Result 1:Performance of different filtering parameters in Samtools fastq:
(1)-f 3: (Output file size:312M)
Choose the flag = 0x1 (paired-end) and the flag = 0x2 (each segment properly aligned according to the aligner) to include the reads needed.
(2)-F 12: (Output file size:317M)
Choose the flag = 4(segment unmapped) and the flag = 8(next segment in the template unmapped) to exclude the reads not needed.
(3)-F 0x900: (Output file size:319M)
Recommended by the help document in order to obtain the only paired reads by excluding the flag = 0x100 (secondary alignment)and the flag = 0x800 (supplementary alignment)
(4)Parameter explainations:
-f: Include the reads with intended flags
-F: Filter the reads with intended flags
-G: Exclude the reads with intended flags
**
Flags explainations:**
Expressed in a hexadecimal or decimal way consists of the following values:
(5)Result: **The lowest assembly contigs could be obtained by extracting subreads with the parameter -f 3**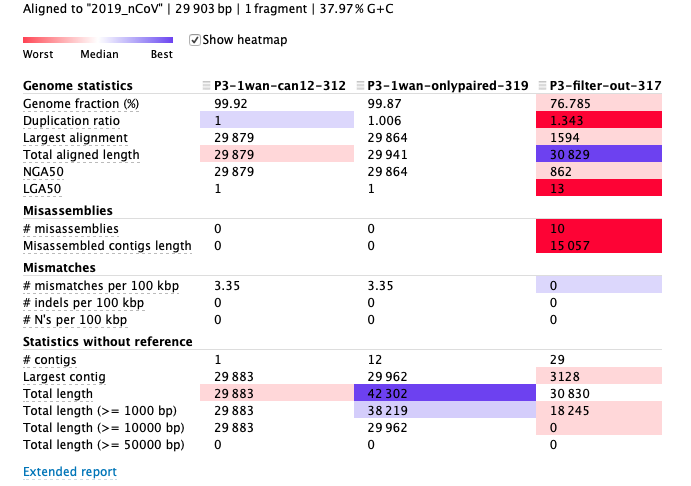
**
Result 2: Assess the assemblies with different subsampling coverage and SPAdes parameters (—meta/—isolate)
30X 100X 1000X 10000X 100000X
The third sample as an example: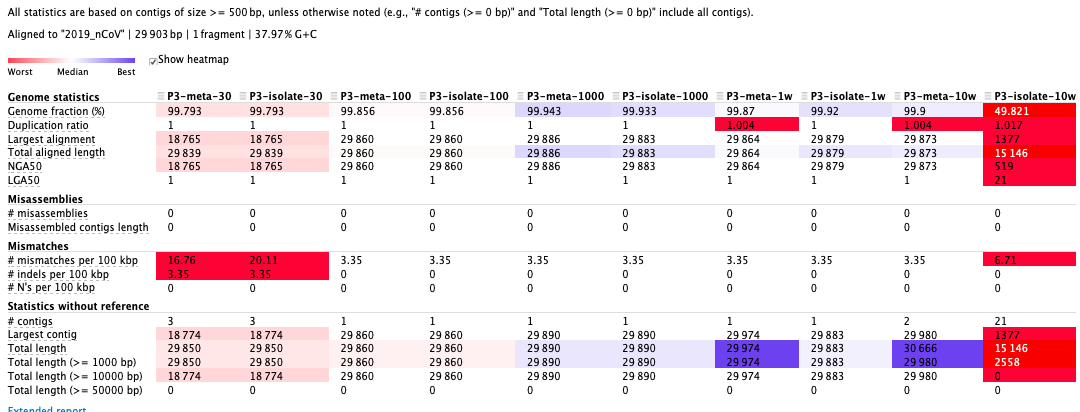
The fifth sample as an example:**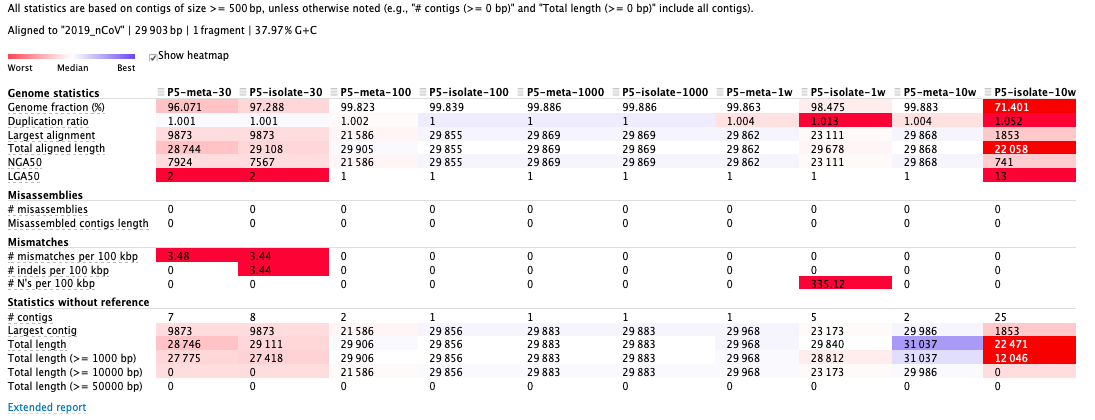
Result:Reads are better assembled with 1000 ~ 10000 subsampling coverage. As for SPAdes parameters, assemblies by “—meta” performed better than “—isolate” with less assembled contigs.
**
Result 3: Package the pipeline :
(1)Goals:
Input the original paired-end fastq files、index files、subsampling fraction and assembling strategies, the output file will be the finally assembled fasta files with only single contigs and the whole process taking around 7 hours for each sample.
Usage:
Run the following shell from the command line:#!/bin/bash#SBATCH -e subfqall.err#SBATCH -o subfqall.out#SBATCH -J subfqall#SBATCH --ntasks-per-node=2#SBATCH -N 1#SBATCH -p sugon#SBATCH -t 120:00:00
python ``ymy.py`` ``--filea`` ./P3-VERO-P3-1-vero_L4_1.fq.gz ``--fileb`` ./P3-VERO-P3-1-vero_L4_2.fq.gz ``--index`` ./2019-nCoV.fasta ``--fraction`` 0.0036 ``--startegy`` meta
(2)—help**: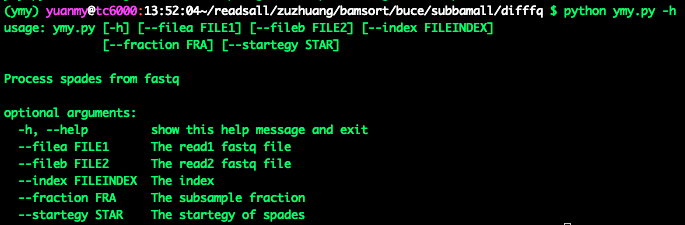
(3)Code:**
#!/usr/bin/python#-*- coding:utf-8 -*-import argparseimport osdef _argparse():parser = argparse.ArgumentParser(description='Process spades from fastq')parser.add_argument('--filea', dest='file1', default='./P3-VERO-P3-1-vero_L4_1.fq.gz', help="The read1 fastq file")parser.add_argument('--fileb', dest='file2', default='./P3-VERO-P3-1-vero_L4_2.fq.gz', help="The read2 fastq file")parser.add_argument('--index', dest='fileindex', default='./2019-nCoV.fasta', help="The index")parser.add_argument('--fraction', dest='fra', default='0.00036', help="The subsample fraction")parser.add_argument('--startegy', dest='star', default='--meta', help="The startegy of spades")return parser.parse_args()def zuzhuang(x,y,z,w,t):'''os.system(f"fastp -i {x} -I {y} -o out1.fastq.gz -O out2.fastq.gz")os.system(f"bwa mem -t 20 {z} out1.fq.gz out2.fq.gz | samtools sort -@ 10 -O bam -o sorted.bam")os.system(f"samtools index -@ 8 sorted.bam sorted.bam.bai")os.system(f"sambamba view -t 10 -h -s {w} sorted.bam -o subsorted.bam")os.system(f"samtools fastq -s -f 3 -1 paired1.fq -2 paired2.fq subsorted.bam")os.system(f"spades.py -{t} -1 paired1.fq -2 paired2.fq -o outputfile")'''os.system("fastp -i {x} -I {y} -o out1.fastq.gz -O out2.fastq.gz".format(x=x, y=y))os.system("bwa mem -t 20 {z} out1.fq.gz out2.fq.gz | samtools sort -@ 10 -O bam -o sorted.bam".format(z = z))os.system("samtools index -@ 8 sorted.bam sorted.bam.bai")os.system("sambamba view -t 10 -h -s {w} sorted.bam -o subsorted.bam".format(w = w))os.system("samtools fastq -s -f 3 -1 paired1.fq -2 paired2.fq subsorted.bam")os.system("spades.py --{t} -1 paired1.fq -2 paired2.fq -o outputfile".format(t = t))#return outputfileprint("all finished!")def main():parser = _argparse()zuzhuang(parser.file1, parser.file2, parser.fileindex, parser.fra, parser.star)if __name__ == '__main__':main()
(4)Performance on 11 samples :
Parameter selection:
Filtering parameter during subsampling : -f 3
Subsampling coverage selection: 1000-10000
SPAdes strategies: —meta,**
Assembled results of 11 samples:
Each sample only has one contig in the assemblie. Some indicators, as the coverage of reference genome, the miss-assembled rates and the error rate are more reliable than before.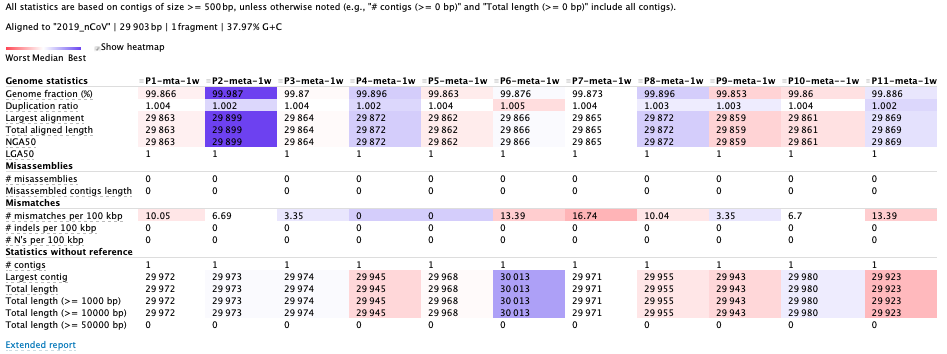
Result 4: Summary:
Several tricky points:
- Ignoring the addition of “-s “ parameter during Samtools fastq can cause errors in subsequent assembly suggesting the reads name are not matched.
- Samtools fastq filtering with “-f 3” parameters can reduce the number of finally assembled contigs to one.
- No improvement is found using the corrected reads from the first SPAdes result files (To note that if you use “—isolate” ,there will be no corrected results. )
- The unpaired reads resulted from SPAdes is useful in Pilon and act as the unpaired bam files.

