1. Introduction:
MAFFT profile
The most classic and widely known multi-sequence alignment software is Clustalw. Comparison speed (Muscle>MAFFT>ClustalW> t-coffee), comparison accuracy (MAFFT>Muscle> t-coffee >ClustalW). Therefore, MAFFT software is recommended for multiple sequence alignment.
2.Detailed software installation and parameters:
The installation
conda install -c bioconda mafft
High speed:
mafft all.fasta > outputmafft --retree 1 in > out (fast)
High accuracy (for <~200 sequences x <~2,000 AA /nt) Three different comparison methods:
mafft --maxiterate 1000 --localpair in > out (% linsi in > out is also ok)
the most accurate method. Suitable for sequences less than 200 and less than 2000AA /nt in length
mafft --maxiterate 1000 --genafpair in > out (% einsi in > out)
The ### is suitable for sequences containing large unmatched regions, less than 200, and less than 2000AA /nt in length
mafft --maxiterate 1000 --globalpair in > out (% ginsi in > out)
Suitable for sequences with similar length alignment, less than 200 and less than 2000AA /nt
If unsure which option to use:
mafft --auto in > out
-- Maxiterate: Maximum number of iteration optimizations, default: 0-- Clustalout: output: Clustal format, default: Fasta-- Reorder: out of order: align, default: input order-Quiet: Don't report progress-- Thread: Number of threads (if not sure, -- Thread-1)
3. Output Results:
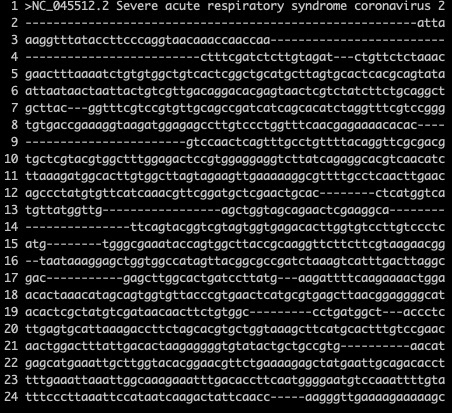
The results are shown in the figure below. According to the longest sequence, mark the aa of the corresponding position. Where there is a gap, fill it out with —.