def loadDataSet():dataMat = []; labelMat = []fr = open('testSet.txt')for line in fr.readlines():lineArr = line.strip().split()dataMat.append([1.0, float(lineArr[0]), float(lineArr[1])])labelMat.append(int(lineArr[2]))return dataMat, labelMatdef sigmoid(inX):return 1.0 / (1 + np.exp(-inX))def gradAscent(dataMatIn, classLabels):dataMatrix = np.mat(dataMatIn) #convert to NumPy matrixlabelMat = np.mat(classLabels).transpose() #convert to NumPy matrixm, n = np.shape(dataMatrix)alpha = 0.001maxCycles = 500weights = np.ones((n, 1))for k in range(maxCycles): #heavy on matrix operationsh = sigmoid(dataMatrix*weights) #matrix multerror = (labelMat - h) #vector subtractionweights = weights + alpha * dataMatrix.transpose()* error #matrix multreturn weights

(获得梯度上升的参数weight,回归系数)
def plotBestFit(weights):import matplotlib.pyplot as pltdataMat, labelMat = loadDataSet()dataArr = np.array(dataMat)n = np.shape(dataArr)[0]xcord1 = []; ycord1 = []xcord2 = []; ycord2 = []for i in range(n):if int(labelMat[i]) == 1:xcord1.append(dataArr[i, 1]); ycord1.append(dataArr[i, 2])else:xcord2.append(dataArr[i, 1]); ycord2.append(dataArr[i, 2])fig = plt.figure()ax = fig.add_subplot(111)ax.scatter(xcord1, ycord1, s=30, c='red', marker='s')ax.scatter(xcord2, ycord2, s=30, c='green')x = np.arange(-3.0, 3.0, 0.1)y = (-weights[0]-weights[1]*x)/weights[2]ax.plot(x, y)plt.xlabel('X1'); plt.ylabel('X2')plt.show()

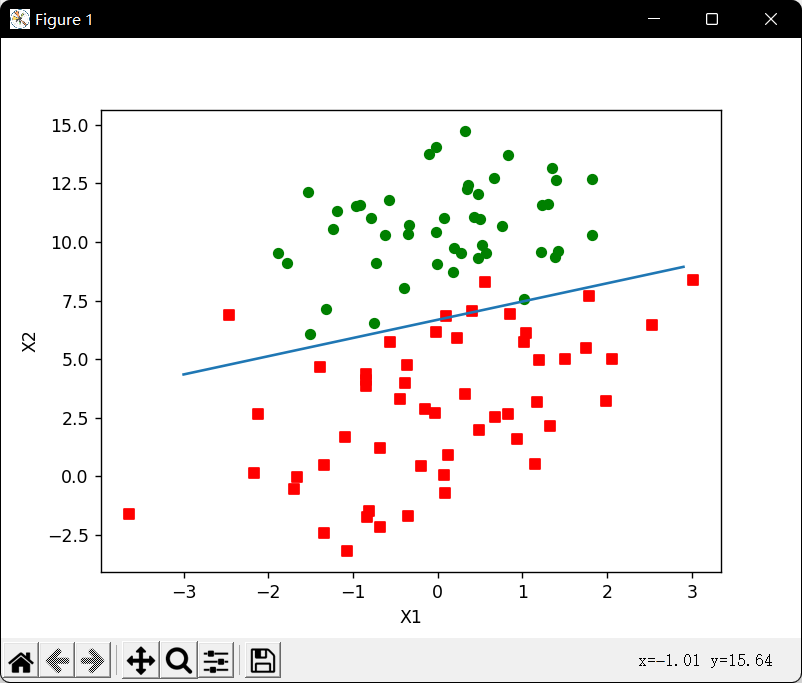
随机梯度上升算法:通过每次用一个样本点来更新回归系数,减少算法复杂度
def stocGradAscent0(dataMatrix, classLabels):m, n = np.shape(dataMatrix)alpha = 0.01weights = np.ones(n) #initialize to all onesfor i in range(m):h = sigmoid(sum(dataMatrix[i]*weights))error = classLabels[i] - hweights = weights + alpha * error * dataMatrix[i]return weights
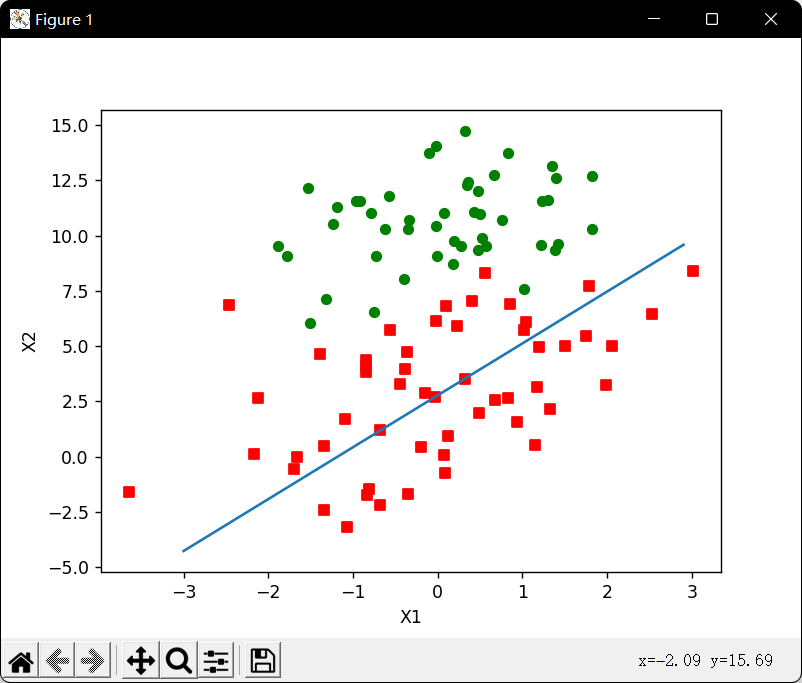

改进随机梯度上升算法:
def stocGradAscent1(dataMatrix, classLabels, numIter=150):m, n = np.shape(dataMatrix)weights = np.ones(n) #initialize to all onesfor j in range(numIter):dataIndex = list(range(m))for i in range(m):alpha = 4/(1.0+j+i)+0.0001 #apha decreases with iteration, does notrandIndex = int(np.random.uniform(0, len(dataIndex)))#go to 0 because of the constanth = sigmoid(sum(dataMatrix[randIndex]*weights))error = classLabels[randIndex] - hweights = weights + alpha * error * dataMatrix[randIndex]del(dataIndex[randIndex])return weights
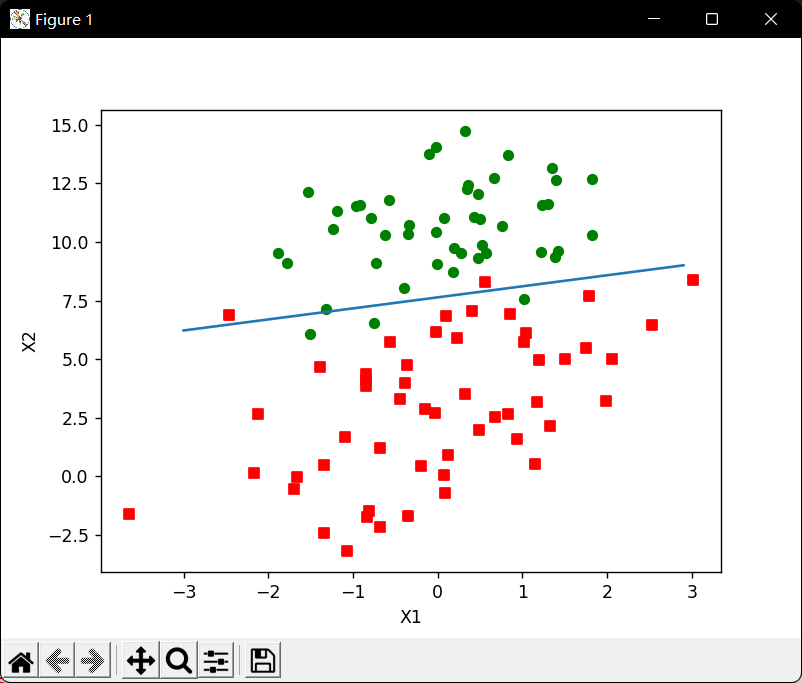

示例:从疝气病预测病马死亡率
def classifyVector(inX, weights):prob = sigmoid(sum(inX*weights))if prob > 0.5: return 1.0else: return 0.0def colicTest():frTrain = open('horseColicTraining.txt'); frTest = open('horseColicTest.txt')trainingSet = []; trainingLabels = []for line in frTrain.readlines():currLine = line.strip().split('\t')lineArr = []for i in range(21):lineArr.append(float(currLine[i]))trainingSet.append(lineArr)trainingLabels.append(float(currLine[21]))trainWeights = stocGradAscent1(np.array(trainingSet), trainingLabels, 1000)errorCount = 0; numTestVec = 0.0for line in frTest.readlines():numTestVec += 1.0currLine = line.strip().split('\t')lineArr = []for i in range(21):lineArr.append(float(currLine[i]))if int(classifyVector(np.array(lineArr), trainWeights)) != int(currLine[21]):errorCount += 1errorRate = (float(errorCount)/numTestVec)print("the error rate of this test is: %f" % errorRate)return errorRatedef multiTest():numTests = 10; errorSum = 0.0for k in range(numTests):errorSum += colicTest()print("after %d iterations the average error rate is: %f" % (numTests, errorSum/float(numTests)))


