SPAdes
SPAdes is an assembly toolkit containing various assembly pipelines. It is mainly used for small genomes, such as bacterial (both single-cell MDA and standard isolates), fungal and other small genomes. SPAdes is not intended for larger genomes (e.g. mammalian size genomes).
options:
--isolate # this flag is highly recommended for high-coverage isolate and multi-cell data--sc # this flag is required for MDA (single-cell) data--meta # this flag is required for metagenomic data-o <output_dir> # directory to store all the resulting files (required)-k <int> [<int> ...] # list of k-mer sizes (must be odd and less than 128)
Setting k-mer as odd is to avoid causing positive and negative chain confusion. SPAdes attempts to assemble genomes using different k-mer sizes, the default list of this parameters is 21 33 55.
Input data:--12 <filename> # file with interlaced forward and reverse paired-end reads-1 <filename> # file with forward paired-end reads-2 <filename> # file with reverse paired-end reads-s <filename> # file with unpaired reads
Assembled genome sequences using SPAdes may be reverse .
Pipeline:
Subsample reads.
$ sambamba view -h -s 0.001 ~/SARS_CoV_2/sortedbam/P3-VERO-P3-9-vero_L4_sort.bam -o ~/SARS_CoV_2/subsample/P3-VERO-P3-9-0.001.bam$ samtools sort -@ 4 -l 9 ~/SARS_CoV_2/subsample/P3-VERO-P3-9-0.001.bam -o ~/SARS_CoV_2/subsample/P3-VERO-P3-9-sort0.001.bam$ samtools index ~/SARS_CoV_2/subsample/P3-VERO-P3-9-sort0.001.bam ~/SARS_CoV_2/subsample/P3-VERO-P3-9-sort0.001.bam.bai
Convert the BAM to FASTQ.
$ samtools fastq ~/SARS_CoV_2/subsample/P3-VERO-P3-9-sort0.001.bam > ~/SARS_CoV_2/fastq/P3-VERO-P3-9_sort0.001.fq
Assemble.
$ spades.py --meta --12 ~/SARS_CoV_2/fastq/P3-VERO-P3-1-vero_L4_006.fq -o ~/SARS_CoV_2/genome/P3-VERO-P3-1-006_meta.fasta
Result:
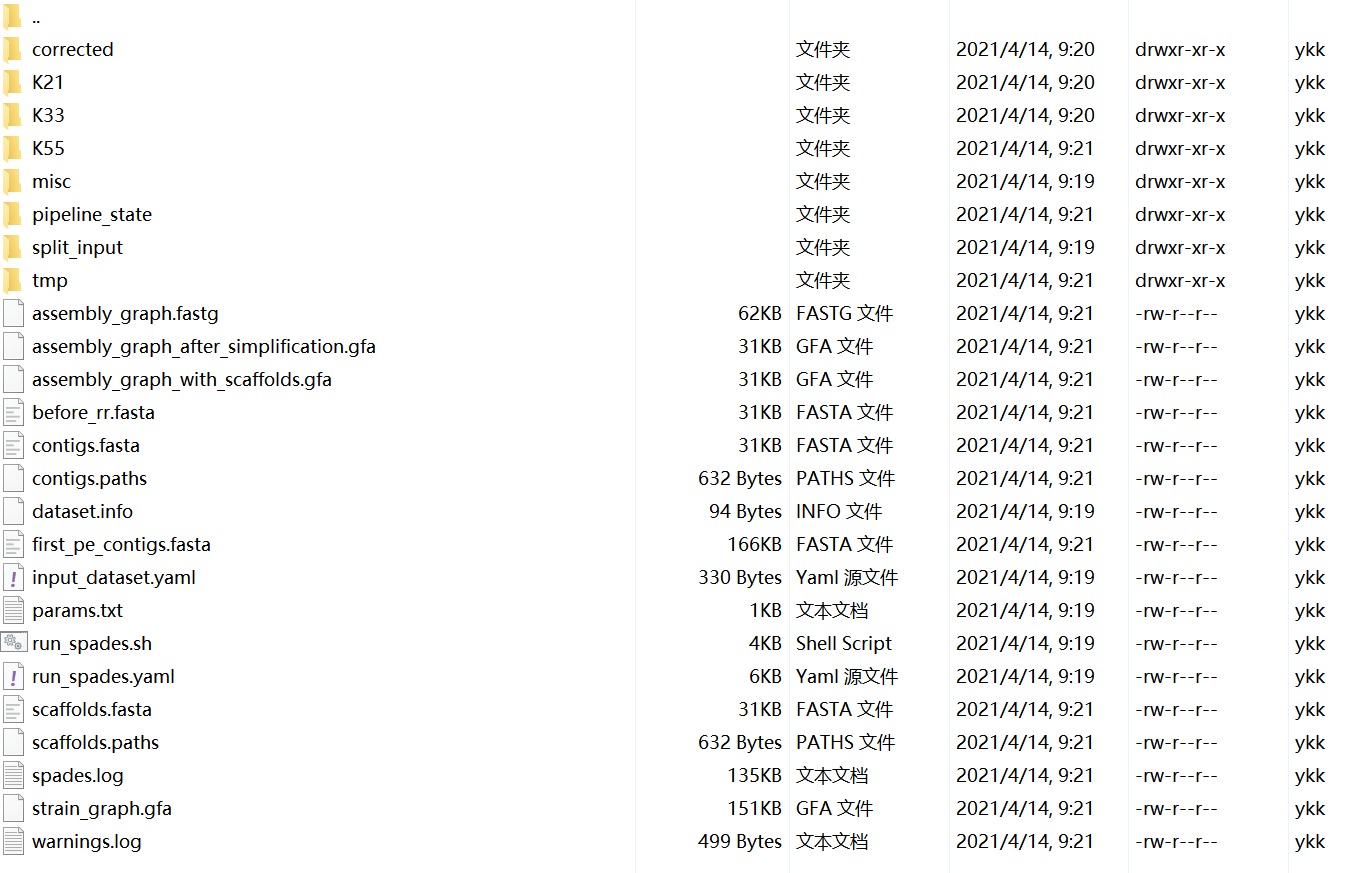
The full list of <output_dir> content is presented below:
scaffolds.fasta– resulting scaffolds (recommended for use as resulting sequences)contigs.fasta– resulting contigs**scaffolds.fasta**and**contigs.fasta**are no difference in the assembly results of this project.assembly_graph.fastg– assembly graphcontigs.paths– contigs paths in the assembly graphscaffolds.paths– scaffolds paths in the assembly graphbefore_rr.fasta– contigs before repeat resolutioncorrected/– files from read error correctionconfigs/– configuration files for read error correctioncorrected.yaml– internal configuration file- Output files with corrected reads
params.txt– information about SPAdes parameters in this runspades.log– SPAdes logdataset.info– internal configuration fileinput_dataset.yaml– internal YAML data set fileK<##>/– directory containing intermediate files from the run with K=<##>. These files should not be used as assembly results; use resulting contigs/scaffolds in files mentioned above.More information:
Invoking the manual
$ conda activate covid$ spades.py

