使用的是来自eggNog的注释信息
目前没有相关的可公开的数据,欢迎有这种数据的小伙伴联系我。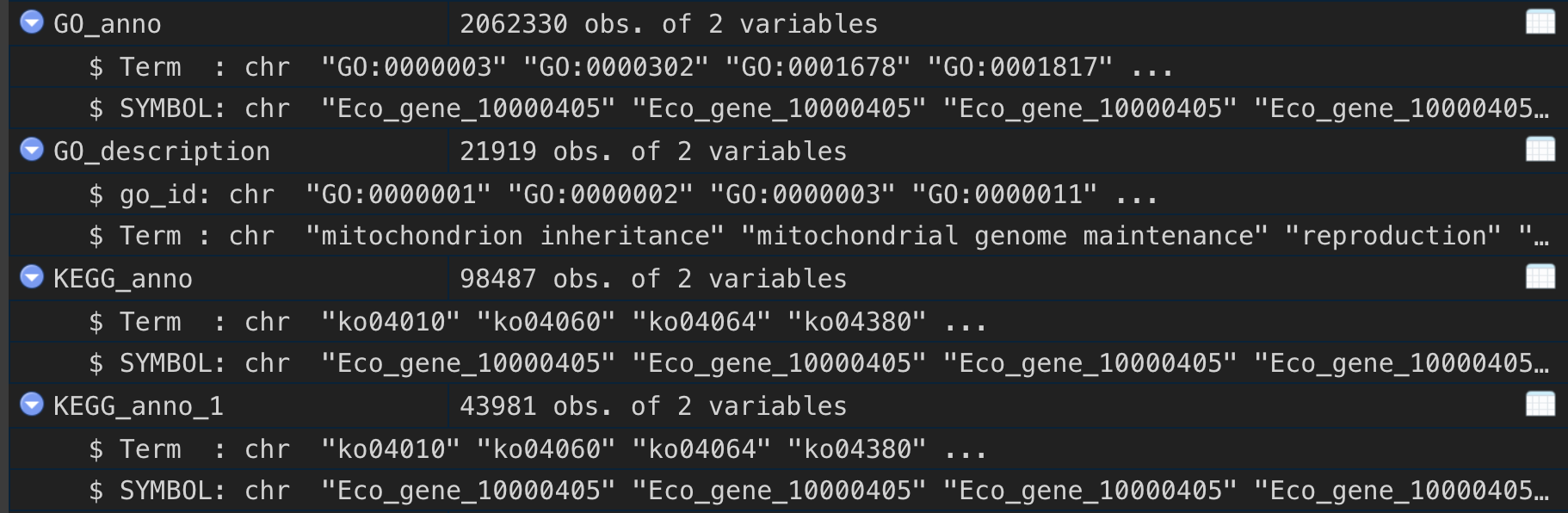
## 安装devtools::install_github("xiayh17/RNAseqStat2")## 载入library(RNAseqStat2)## 读取count矩阵row_counts <- read.delim("count_data.txt", row.names=1)## 分组信息group_list <- c("C","C","C","I","I","I")## 读取注释数据anno_data <- data.table::fread("out.emapper.annotations",skip = 4,fill = T)## GOGO_anno <- eggnogG2T(anno_data,"#query","GOs")## 获取GO Term的描述信息GO_description <- clusterProfiler::go2term(GO_anno$Term)## KEGGKEGG_anno <- eggnogG2T(anno_data,"#query","KEGG_Pathway")KEGG_anno <- KEGG_anno %>%filter(stringr::str_detect(Term, 'ko'))## 获取KEGG的描述信息KEGG_description <- clusterProfiler::ko2name(KEGG_anno$Term)data_i <- Create_DEGContainer(expMatrix = row_counts,groupInfo = group_list,caseGroup = "C",idType = "SYMBOL",species = "unknow",GOTERM2GENE = GO_anno,GOTERM2NAME = GO_description,KEGGTERM2GENE = KEGG_anno,KEGGTERM2NAME = KEGG_description)## 一键全部data_o <- runALL(object = data_i,dir = "output_test",GO = TRUE,KEGG = TRUE)## 分步运行runCheck(data_i,"test_check")data_g <- runDEG(obj = data_i, parallel = T,dir = "test_deg")#data_h <- runHyper(obj = data_g,dir = "test_hyper",GO = TRUE,KEGG = TRUE)data_gse <- runGSEA(obj = data_h,dir = "test_gse",GO = TRUE,KEGG=TRUE)

