语雀:左手柳叶刀右手炭火烧
微信公众号:研平方 | 简书:研平方
关注可了解更多的科研教程及技巧。如有问题或建议,请留言。
欢迎关注我:一起学习,一起进步!
最近,小编“扫荡”文献时,发现一个令我十分感兴趣的应用,利用文本文本挖掘技术可以评估选定基因与癌症之间的关联。提到文本挖掘这类技术,小编当然要一探究竟了。
1.原文如下
Literature evidence for the identified target genes in cancer
We used OncoScore, a text mining tool to assess the associations between each gene and specific cancers based on the literature. A cutoff value of 21.09 was suggested to determine true positives and the true negatives in cancer gene identification.
2.查找资料
习惯性的打开浏览器,准备打破砂锅问到底,惊喜的发现,OncoScore竟然是一个写好的R包,而且放在了Bioconductor网页,可直接进行安装、使用。虽然文章发在了Sci Rep杂志上,但是小编认为还是值得一试。
3.它能干什么
The OncoScore analysis consists of two parts. One can estimate a score to asses the
oncogenic potential of a set of genes, given the lecterature knowledge, at the time of the
analysis, or one can study the trend of such score over time.
可见,OncoScore不仅可以依据文献中的知识,对一组设定目标基因列表的致癌能力进行评分,还可以研究这个分数随时间的趋势。
4.开始表演,拿好小板凳看戏
4.1 准备工作
if (!requireNamespace("BiocManager", quietly = TRUE))install.packages("BiocManager")BiocManager::install("OncoScore")# load the librarylibrary(OncoScore)# Define a queryquery = perform.query(c("ASXL1","IDH1","IDH2","SETBP1","TET2"))### Starting the queries for the selected genes.### Performing queries for cancer literatureNumber of papers found in PubMed for ASXL1 was: 923Number of papers found in PubMed for IDH1 was: 3691Number of papers found in PubMed for IDH2 was: 1318Number of papers found in PubMed for SETBP1 was: 177Number of papers found in PubMed for TET2 was: 1609### Performing queries for all the literatureNumber of papers found in PubMed for ASXL1 was: 1018Number of papers found in PubMed for IDH1 was: 3902Number of papers found in PubMed for IDH2 was: 1499Number of papers found in PubMed for SETBP1 was: 229Number of papers found in PubMed for TET2 was: 2117
以上我们可以发现,通过检索,得到了癌症相关研究的文献数量,以及所有与检索基因相关文献数量。
OncoScore provides a function to merge gene names if requested by the user. This function is useful when there are aliases in the gene list.
combine.query.results(query, c('IDH1', 'IDH2'), 'new_gene')CitationsGene CitationsGeneInCancerASXL1 1018 923SETBP1 229 177TET2 2117 1609new_gene 5401 5009
当然,OncoScore还可以依据染色体信息检索基因。这里不再演示。
4.2 重点来啦
4.2.1 开始计算基因的致癌评分
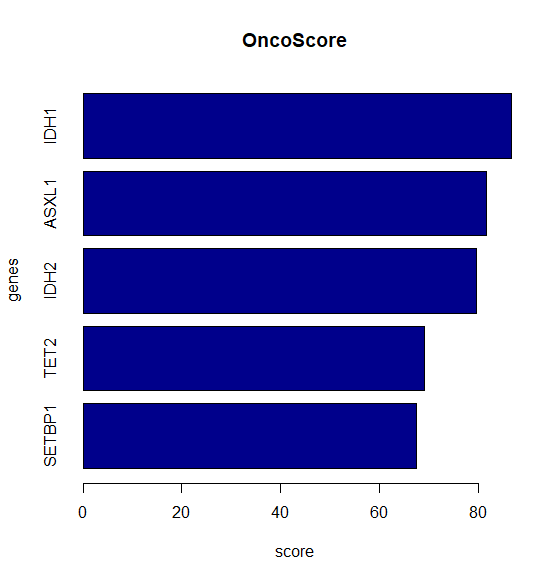
result = compute.oncoscore(query)### Processing data### Computing frequencies scores### Estimating oncogenes### Results:ASXL1 -> 81.59349IDH1 -> 86.66355IDH2 -> 79.59096SETBP1 -> 67.43283TET2 -> 69.12424
4.2.2 时间趋势分析(OncoScore timeline analysis)
query.timepoints = perform.query.timeseries(c("ASXL1","IDH1","IDH2","SETBP1","TET2"),c("2012/03/01", "2013/03/01", "2014/03/01", "2015/03/01", "2016/03/01"))### Starting the queries for the selected genes.### Quering PubMed for timepoint 2012/03/01### Performing queries for cancer literatureNumber of papers found in PubMed for ASXL1 was: 86Number of papers found in PubMed for IDH1 was: 409Number of papers found in PubMed for IDH2 was: 173Number of papers found in PubMed for SETBP1 was: 5Number of papers found in PubMed for TET2 was: 173### Performing queries for all the literatureNumber of papers found in PubMed for ASXL1 was: 92Number of papers found in PubMed for IDH1 was: 489Number of papers found in PubMed for IDH2 was: 235Number of papers found in PubMed for SETBP1 was: 10Number of papers found in PubMed for TET2 was: 197### Quering PubMed for timepoint 2013/03/01### Performing queries for cancer literatureNumber of papers found in PubMed for ASXL1 was: 135Number of papers found in PubMed for IDH1 was: 662Number of papers found in PubMed for IDH2 was: 267Number of papers found in PubMed for SETBP1 was: 11Number of papers found in PubMed for TET2 was: 258### Performing queries for all the literatureNumber of papers found in PubMed for ASXL1 was: 150Number of papers found in PubMed for IDH1 was: 753Number of papers found in PubMed for IDH2 was: 336Number of papers found in PubMed for SETBP1 was: 18Number of papers found in PubMed for TET2 was: 303### Quering PubMed for timepoint 2014/03/01### Performing queries for cancer literatureNumber of papers found in PubMed for ASXL1 was: 188Number of papers found in PubMed for IDH1 was: 904Number of papers found in PubMed for IDH2 was: 365Number of papers found in PubMed for SETBP1 was: 29Number of papers found in PubMed for TET2 was: 347### Performing queries for all the literatureNumber of papers found in PubMed for ASXL1 was: 209Number of papers found in PubMed for IDH1 was: 1003Number of papers found in PubMed for IDH2 was: 440Number of papers found in PubMed for SETBP1 was: 36Number of papers found in PubMed for TET2 was: 431### Quering PubMed for timepoint 2015/03/01### Performing queries for cancer literatureNumber of papers found in PubMed for ASXL1 was: 257Number of papers found in PubMed for IDH1 was: 1198Number of papers found in PubMed for IDH2 was: 468Number of papers found in PubMed for SETBP1 was: 51Number of papers found in PubMed for TET2 was: 461### Performing queries for all the literatureNumber of papers found in PubMed for ASXL1 was: 286Number of papers found in PubMed for IDH1 was: 1304Number of papers found in PubMed for IDH2 was: 551Number of papers found in PubMed for SETBP1 was: 66Number of papers found in PubMed for TET2 was: 583### Quering PubMed for timepoint 2016/03/01### Performing queries for cancer literatureNumber of papers found in PubMed for ASXL1 was: 323Number of papers found in PubMed for IDH1 was: 1506Number of papers found in PubMed for IDH2 was: 569Number of papers found in PubMed for SETBP1 was: 68Number of papers found in PubMed for TET2 was: 587### Performing queries for all the literatureNumber of papers found in PubMed for ASXL1 was: 359Number of papers found in PubMed for IDH1 was: 1625Number of papers found in PubMed for IDH2 was: 661Number of papers found in PubMed for SETBP1 was: 89Number of papers found in PubMed for TET2 was: 745
perform.query.timeseries ()函数检索了几个设定时间的文献数据信息。
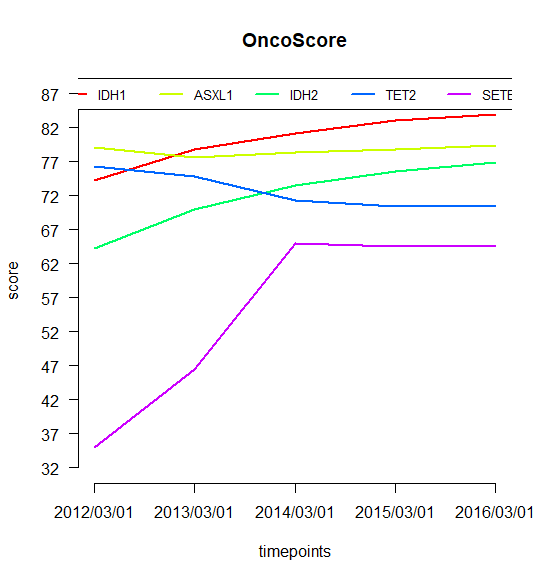
result.timeseries = compute.oncoscore.timeseries(query.timepoints)### Computing oncoscore for timepoint 2012/03/01### Processing data### Computing frequencies scores### Estimating oncogenes### Results:ASXL1 -> 79.14893IDH1 -> 74.27776IDH2 -> 64.27063SETBP1 -> 34.9485TET2 -> 76.29579### Computing oncoscore for timepoint 2013/03/01### Processing data### Computing frequencies scores### Estimating oncogenes### Results:ASXL1 -> 77.54983IDH1 -> 78.71551IDH2 -> 69.99559SETBP1 -> 46.4559TET2 -> 74.81894### Computing oncoscore for timepoint 2014/03/01### Processing data### Computing frequencies scores### Estimating oncogenes### Results:ASXL1 -> 78.28121IDH1 -> 81.08963IDH2 -> 73.50788SETBP1 -> 64.97398TET2 -> 71.31087### Computing oncoscore for timepoint 2015/03/01### Processing data### Computing frequencies scores### Estimating oncogenes### Results:ASXL1 -> 78.84769IDH1 -> 82.99363IDH2 -> 75.60886SETBP1 -> 64.48853TET2 -> 70.46695### Computing oncoscore for timepoint 2016/03/01### Processing data### Computing frequencies scores### Estimating oncogenes### Results:ASXL1 -> 79.37202IDH1 -> 83.9881IDH2 -> 76.89328SETBP1 -> 64.60591TET2 -> 70.53378
4.2.3 可视化
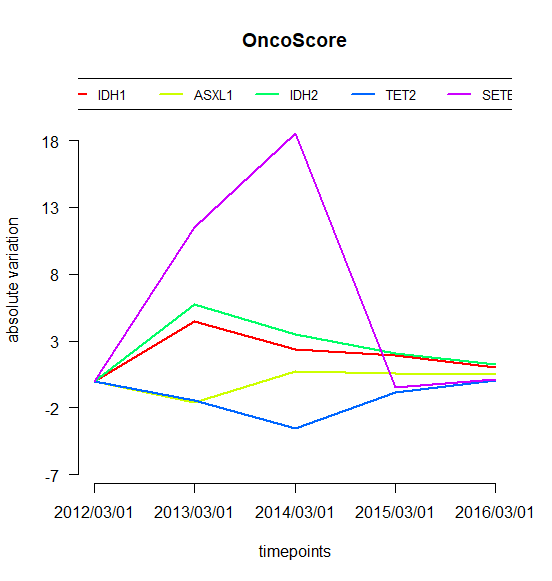
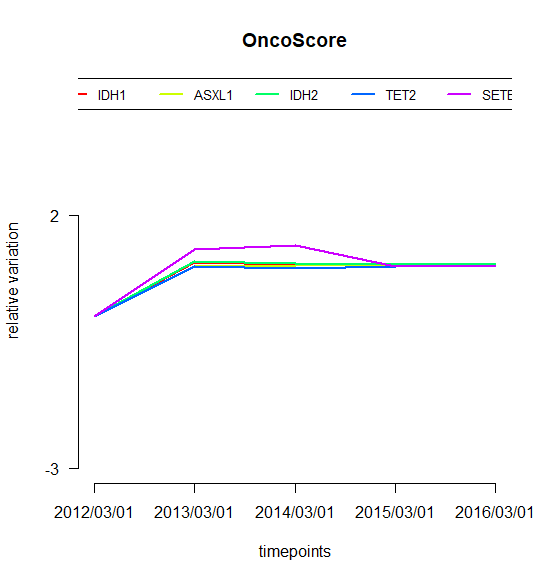
## Oncogenetic potential of the considered genesplot.oncoscore(result, col = 'darkblue')## Absolute values of the oncogenetic potential of the considered genes over timesplot.oncoscore.timeseries(result.timeseries)## Variations of the oncogenetic potential of the considered genes over timesplot.oncoscore.timeseries(result.timeseries,incremental = TRUE,ylab='absolute variation')## Variations as relative values of the oncogenetic potential of the considered genes over timesplot.oncoscore.timeseries(result.timeseries,incremental = TRUE,relative = TRUE,ylab='relative variation')